During my summer internship for the Biodesign project, I went to Pr. Van der Meer’s lab.
Pr. Van der Meer’s lab is interested on understanding microbial activities in the environment at a fundamental level, how microbial communities are influenced by human activities and how they can apply microbial processes for improving the quality of our living environment.
In the context of our project, this lab provides us the GMO bacteria which are able to detect arsenic, and also gives us the feedback about the bioreporter part.
The aim of this week was to learn more about microbiology and thus how the bacteria we use for arsenic detection are designed.
That is why I have performed many experiments to clone some bacteria, like making competent cells, PCR, restriction digestion… It was not bacteria able to detect arsenic, but bacteria in which we studied how the ribose fixation takes place. We cultivated some bacteria with modified genome and we saw what is the consequence in the ribose pathway. But the principle of cloning stays the same and it was very interesting to understand how the bioreporter is made.
Pr. Van der Meer and his PhD student, Vitali Maffenbeier, were also interested about the prototype 1, to measure the fluorescence emited by the bioreporters.
Thus we have prepared some bacteria which produces eGFP in presence of ribose, as model system. We added to our overnight culture of bacteria either 10 or 100 ug/L of ribose. After 6h of incubation, we made a measurement with our device and with a fluorimeter.
Aim of the experiment : study of different populations of bacteria, which have each a mutation in the coding region of the transmembrane transporter (coupled with eGFP) involved in the ribose pathway and determine which mutation has a negative impact in this pathway thanks to the fluorescence measurement.
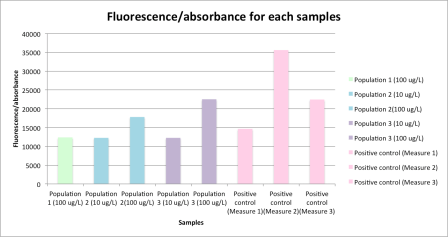
Here are the results we get :
Figure 1 – Fluorescence/absorbance measured with the prototype 1 for each samples
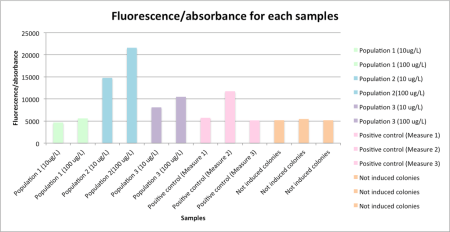
Figure 2 – Fluorescence/absorbance measured with the lab fluorimeter for each samples
As expected, we get a lower fluorescence for the not induced colonies, our negative control. Moreover, we have more fluorescence for the sample with a final concentration of ribose of 200 ug/L than with 20 ug/L, especially for the population 2 and 3. However for the population 1, the fluorescence stays similar to the negative control fluorescence. It shows that there is a problem with the ribose transporter production, which is coupled with eGFP production. Finally, it seems we have a problem with our positive control, which must present a high level of fluorescence (like for the measure 2).
In this study, we determine that the population 1, in which we measure a fluorescence similar to the fluorescence of the negative control, presents a fatal mutation in the ribose transporter coding sequence.